Fu, L.; Jung, Y.; Tian, C.; Ferreira, R.; He, F.; Yang, J.; Carroll, K. ChemRxiv 2022.
https://chemrxiv.org/engage/chemrxiv/article-details/62ab096604a3a9469c48d4ec
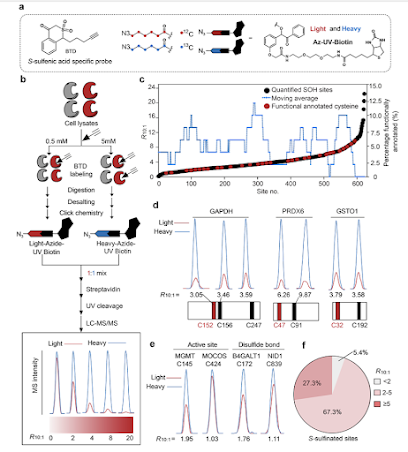
The convergence of reactive cysteine-targeted electrophilic fragments and chemoproteomics have dramatically accelerated the discovery of ligandable sites in the proteome. Our genome encodes 214,000 cysteine residues, at least 20% of which are estimated to be redox-active. Oxidation blunts sulfur reactivity toward electrophiles but opens the door to a new class of nucleophilic covalent ligands that target cysteinyl sulfenic acids, which are widespread post-translational modifications. Here we report a quantitative analysis of nucleophilic fragments screened against the human sulfenome. Ligands were discovered for >500 sulfenated cysteines in >400 proteins, including sites not targeted by electrophiles with the same scaffold. Among these were compounds that preferentially react with hepatoma-derived growth factor (HDGF)-related proteins (HRPs) one of which was able to block nuclear transport of this oncoprotein. Nucleophilic fragments provide a rich resource for chemical biology and drug discovery, where ligandability in the human proteome extends beyond protein thiols.